Scaling Down: The Python Libraries You Need to Compress and Analyze the PUMS Dataset
By
K
Kae SuarezandM
Marlene MhangamiAuthor(s):
K
Kae SuarezM
Marlene MhangamiPublished: March 7, 2023
13 min read
TL;DR
you can import 14GB of CSV into Parquet blindly, but we can do better.
The Dataset: What is PUMS?
The PUMS (Public Use Microdata Sample) census dataset is a detailed collection of information on a sample of the population, including demographic, housing, and economic characteristics. It is collected by the U.S. Census Bureau as part of the decennial census and is made available to the public for research and analysis. However, due to its large size, analyzing the PUMS dataset is difficult. Today, we will show you how it is possible to wrangle it with just a couple of tools.
What do we want to do?
The dataset is useful for a wide range of stakeholders, including researchers, policymakers, and businesses, as it provides a detailed picture of the population which can inform decision-making and policy-making. Today, we’ll explore one five-year window — the dataset is published regularly with new features and entries, year-by-year. At 3.1 GB compressed, 14 GB uncompressed, and 50GB in memory, almost a hundred files and 200+ columns, wading through it requires a fairly large computer or perhaps access to a cloud-based tool. And after loading the entire dataset, any slicing or dicing can bring most tools to a crawl. This type of tabular data could, of course, be broken down to subsets that are loaded one at a time, but in this article, we will show how a one-time transformation of the entire dataset into a Parquet format can make it compact, fast to use, and portable.
How do we accomplish this? First, we need the data.
Getting the Data
The U.S. Census provides the PUMS data in several ways. You can browse the data right on their website. But after selecting 10 variables (out of 522) we are met with a throttle limit on how much data we can select at a time.
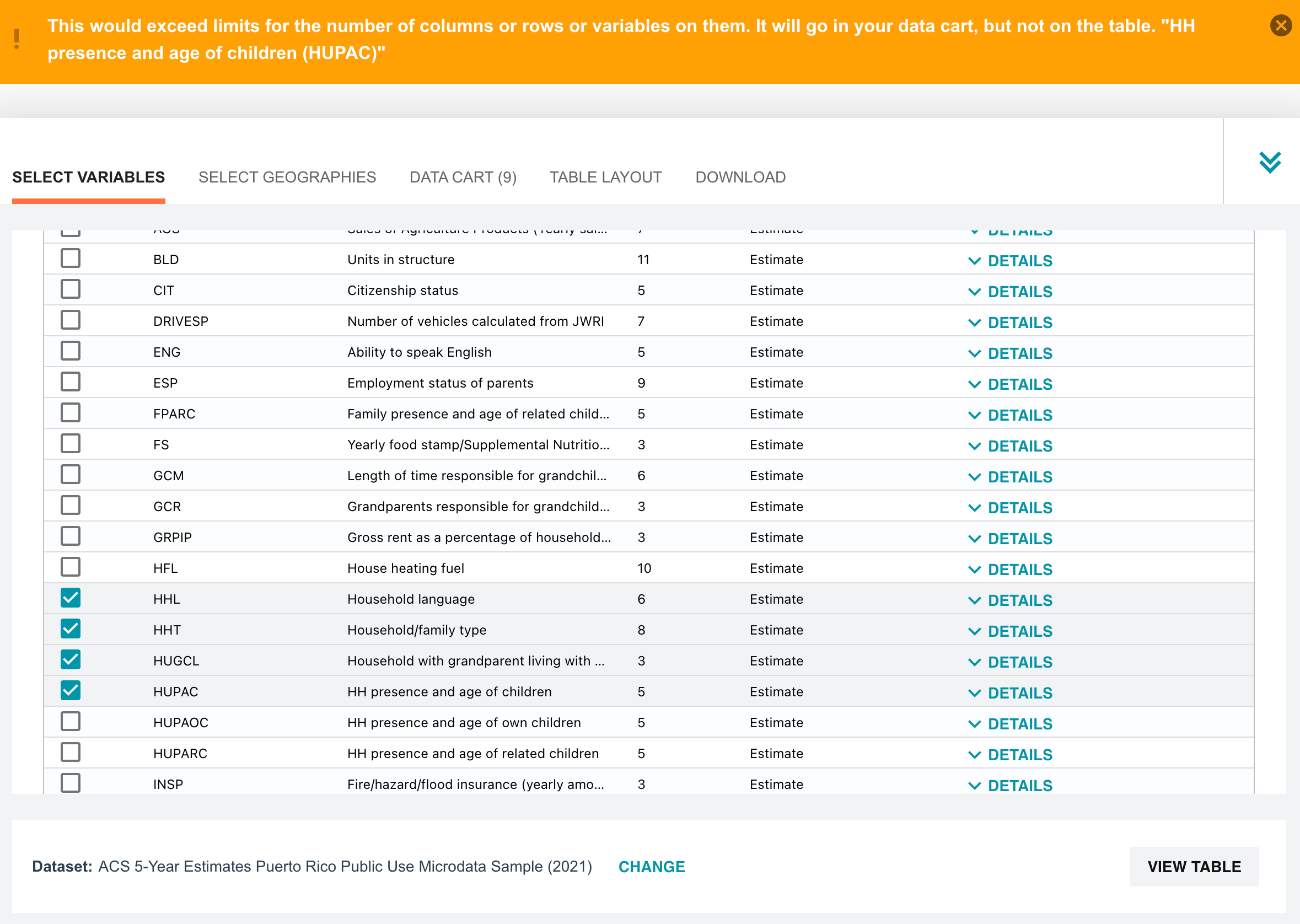
But lucky for us, census.gov also provides data via their HTTP and FTP servers. We chose the five-year data from 2020, found here, and downloaded it in CSV format [HTTP link] and FTP link: ftp://ftp.census.gov/programs-surveys/acs/data/pums/2020/5-Year/. From there, use your preferred tool, such as wget or aria2.
From CSV to Parquet
💡 Note: We are using Jupyter notebooks. The code can be made available upon request.
Now that we have the files, we can get to work. The data we just downloaded is in the form of a large set of CSV’s. We’ll transform this group of files to a single Parquet file. Other than Jupyter, we’re going to use Ibis and PyArrow, so make sure to have those installed with your Python package manager of choice before trying this out!
Once on disk, there is some small housekeeping to do before we can use the data. First, it is worthwhile to discard the csv_pus.zip and csv_hus.zip files, as these contain full-US data, with the exception of Puerto Rico — while the remainder of the files contains details on each state, including Puerto Rico. We will use the latter collection.
After unzipping the files, we’ll divide the five-year set into person-focused and housing-focused data. Person data can be identified by the naming pattern psam_p*.csv, while housing data has the pattern psam_h*.csv. From there, you can use globbing to move them where you want them — the example below will work on Mac and Linux.
bash
mkdir housing
mkdir people
mv psam_h* ./housing/
mv psam_p* ./people/Once this is done, you should now have two directories full of CSV data that make up approximately 14.13 GB.
14 GB is far too big for a desktop environment, and there are too many files in each directory to wrangle efficiently. Not to mention that, if you were to read the values directly, you may see them balloon to 50 GB manifested in memory if your tool prefers 64-bit integers. Once we convert the CSV’s to just two Parquet files (one for household, and one for person data), the data will sit at 4 GB, but with all the features of Parquet, such as embedded metadata and columnar formatting!
Method 1: Quick but Naive
To start, we’ll use the most direct route to convert the CSV files to Parquet. Reading in all of the CSV files at once could overwhelm the system memory, but Apache Arrow can handle that natively. We’re going to use Python today, so we’ll specifically use PyArrow.
If you’re following along, make one notebook for housing data, and one for people data, or just pick one. The same code works for both, except for one line in this section, which we call out. We chose not to combine the datasets, since doing so felt against the spirit of the original dataset.
We have some specific dependencies we’ll need, so the first step is getting those imported:
python
import pyarrow as pa
from pyarrow import parquet
import pyarrow.dataset as dsNow, we need a view of the data — we’ll use PyArrow’s dataset functionality to do this. Note that we exclude invalid files. This is a file format-specific check and ensures we don’t ingest any other files, such as the notebooks themselves.
python
# Open dataset with new schema
dataset = ds.dataset('.', format="csv", exclude_invalid_files=True)Due to a quirk of the dataset, it’s very possible for inference to read a couple of variables as numeric when they are strings in later rows. To handle this, we redefine the schema and make a new view. This is a cheap operation because we haven’t read any data yet, only set up a view that’s aware of the files.
python
# First line needed for person and housing data, second for person only.
nuschem = dataset.schema.set(1, pa.field("SERIALNO", pa.string()))
nuschem = nuschem.set(75, pa.field("WKWN", pa.string()))
#Read using the new schema.
dataset = ds.dataset('.', format="csv", exclude_invalid_files=True, schema=nuschem)Now, we write to Parquet. This can take a while, as it’s only now that PyArrow fully reads the data.
python
ds.write_dataset(dataset, "naive_pums", format="parquet")What gains do we get from this? Well, for person data, the CSVs make up around 10 GB, and the Parquet file occupies 3.3 GB. Household CSVs, of course, make up the other 4 GB of CSV files. If we apply this process to household data, that Parquet is 1.3 GB.
This may feel unfair since Parquet is compressed — and, if we compare to the compressed data, these Parquet files are larger than the original zip files, which were 3.1 GB for all of the CSV. However, Parquet versions are packed into individual files that can be read directly, instead of having to unzip all the CSV data and manifest all 14 GB on disk.
Earlier, it was mentioned that stronger type inference would be important, and this is a great time to see why. Writing to Parquet used the inferred schema from before, and when we read that into memory and query the size, like so:
python
parquet.read_table("./naive_pums/part-0.parquet").get_total_buffer_size()The output is 36432055629 or 36 GB, just for person data — adding household data brings us to 50 GB overall. How did this happen? PyArrow’s type inference tries to be safe when reading in large datasets — we don’t want to read the whole file, so it makes the safest guess, which is that anything numeric will need a 64-bit signed integer. Being wrong has the consequence of misrepresenting data or failing entirely, so it makes sense to do things safely.
Method 2: Analytics for Type Certainty
Reviewing the data shows that a great deal of it needs 32-bits at most, and some are able to be boolean, as the column can only have 0, 1, or blank. So, using a 64-bit integer for every number is too safe for our dataset, but we can’t expect PyArrow to just know that. In many cases, this is fine — but we have a lot of data, and want to make a one-time change that can improve future executions. We have time and memory to spare to do better. We can do some analytics to make something stronger.
This is where we turn to Ibis. PyArrow is a strong tool for everything it does, but Ibis is a clear choice for data analytics. Ibis is an interface that allows easy and standard analytics, defined in Ibis and executed on one of its 15+ backends, ranging from Polars to BigQuery. Here, we get to stay in memory (give or take some swap space), so let’s use one of the many backends Ibis offers, which performs well in memory: DuckDB.
Analytics with Ibis and DuckDB
Let’s do what we did above, and start with imports.
python
import ibis
from ibis import selectors as s
import pyarrow as pa
from pyarrow import parquet
import pyarrow.dataset as dsDuckDB’s the default, but for the sake of certainty, we’ll set it and read with it explicitly.
python
ibis.set_backend("duckdb")
t = ibis.read_csv("psam_p*.csv", sample_size = -1)We’re going to use PyArrow for writing to our new optimized table because it’s so good at that — thus, when building up the schema, we’re going to use PyArrow types. This function is a very brute-force way to do that, which should still be quick since we’re going to use it on an amount of data equal to the number of columns, rather than row*columns.
python
def vals_to_type(minimum, maximum):
if minimum < 0:
if -128 <= minimum <= 127 and -128 <= maximum <= 127:
return pa.int8()
if -32768 <= minimum <= 32767 and -32768 <= maximum <= 32767:
return pa.int16()
if -2147483648 <= minimum <= 2147483647 and -2147483648 <= maximum <= 2147483647:
return pa.int32()
else:
if 0 <= minimum <= 1 and 0 <= maximum <= 1:
return pa.bool_()
if 0 <= minimum <= 255 and 0 <= maximum <= 255:
return pa.uint8()
if 0 <= minimum <= 65535 and 0 <= maximum <= 65535:
return pa.uint16()
if 0 <= minimum <= 4294967295 and 0 <= maximum <= 4294967295:
return pa.uint32()
return pa.int64()There’s no handling for floating point values or non-numbers in general — our dataset is of a form that allows us to focus on integers, and we can handle strings before we find minimum and maximum values.
As for how we set up for our minimum and maximum values, and exclude strings, we can use selectors from Ibis:
python
aggs = []
num_cols = t.select(s.numeric()).schema().names
str_cols = t.select(s.of_type(ibis.expr.datatypes.string)).schema().names
for col in num_cols:
aggs.append(t[col].min())
aggs.append(t[col].max())Now, it may look like we’ve made an array out of results, but this is actually a touch more interesting. Running through getting min then max for each result could take a very, very long time — and optimizers exist to make this better. Ibis gives us a path to gather several results we want and set up the necessary SQL to make it happen, via its concept of expressions. Rather than an array of results, we have an array of unexecuted expressions. We can pass this array of expressions to Ibis and it will set up to run them as smartly as possible in the backend.
Here’s how we do that — on an M2 Macbook Pro, this takes around 1 minute, 40 seconds over the whole dataset:
python
# To break this down: t is the table, .aggregate() is what can
# take a set of expressions, and .execute() actually forces the
# aggregated expression set to run and output -- to a pandas Dataframe
# by default.
out = t.aggregate(aggs).execute()After running that cell, out looks something like this:
python
Min(PUMA) Max(PUMA) Min(REGION) Max(REGION) Min(ST) Max(ST) ...
0 9 1 20 100 70301 . . .
1 rows × 568 columnsUsing Analytics for a Better Schema
Now, we want to set up for PyArrow to use this as its schema — PyArrow can accept a dictionary, so we’ll use the columns’ names as keys, and assign a data type to use as the value. That looks like this:
python
# Set up to iterate over the Pandas Dataframe
test_iter = iter(out.items())
idx = 0
dict_fields = {}
for item in test_iter:
# For each min-max pair in the Dataframe, use vals_to_type() to find
# the appropriate data type using the current loop iteration (min),
# and skip to the next (max).
dict_fields[num_cols[idx]] = vals_to_type(item[1].values[0], next(test_iter)[1].values[0])
# We are guaranteed to have the right order, so we can just increment
# an index value over the numeric columns.
idx += 1
# The string columns are simple -- we just need to make each one map to
# string.
for col in str_cols:
dict_fields[col] = pa.string()PyArrow expects the schema to be in order when passed for reading. This dictionary is not in order. However, we can use the trick from the naive method, in which we make a view that establishes the initial schema, then modify it.
python
dataset = ds.dataset('.', format="csv", exclude_invalid_files=True)
# We have to build the schema out of place, so we copy the
# original to get position data, then set up a schema based on that.
nuschem = dataset.schema
for field in dict_fields.items():
nuschem = nuschem.set(nuschem.get_field_index(field[0]), pa.field(field[0], field[1]))
# With a properly ordered version of our engineered schema, we can now
# make a new view.
dataset = ds.dataset('.', format="csv", exclude_invalid_files=True, schema=nuschem)Results
Now, we have a dataset with our new and improved schema. We’ll write it, just like in the naive method:
python
ds.write_dataset(dataset, "brute_schema_pums", format="parquet")On disk, this yields…a 3 GB file for person data, and 1.2 GB for household data. Seems like the compression is what offers most of the gains — doing this work for a better schema is 10% better than a Parquet without this, and, in comparison to the CSV dataset, this is only a 3% change. Disappointing, but what’s it look like in memory?
python
parquet.read_table("./brute_schema_pums/part-0.parquet").get_total_buffer_size()
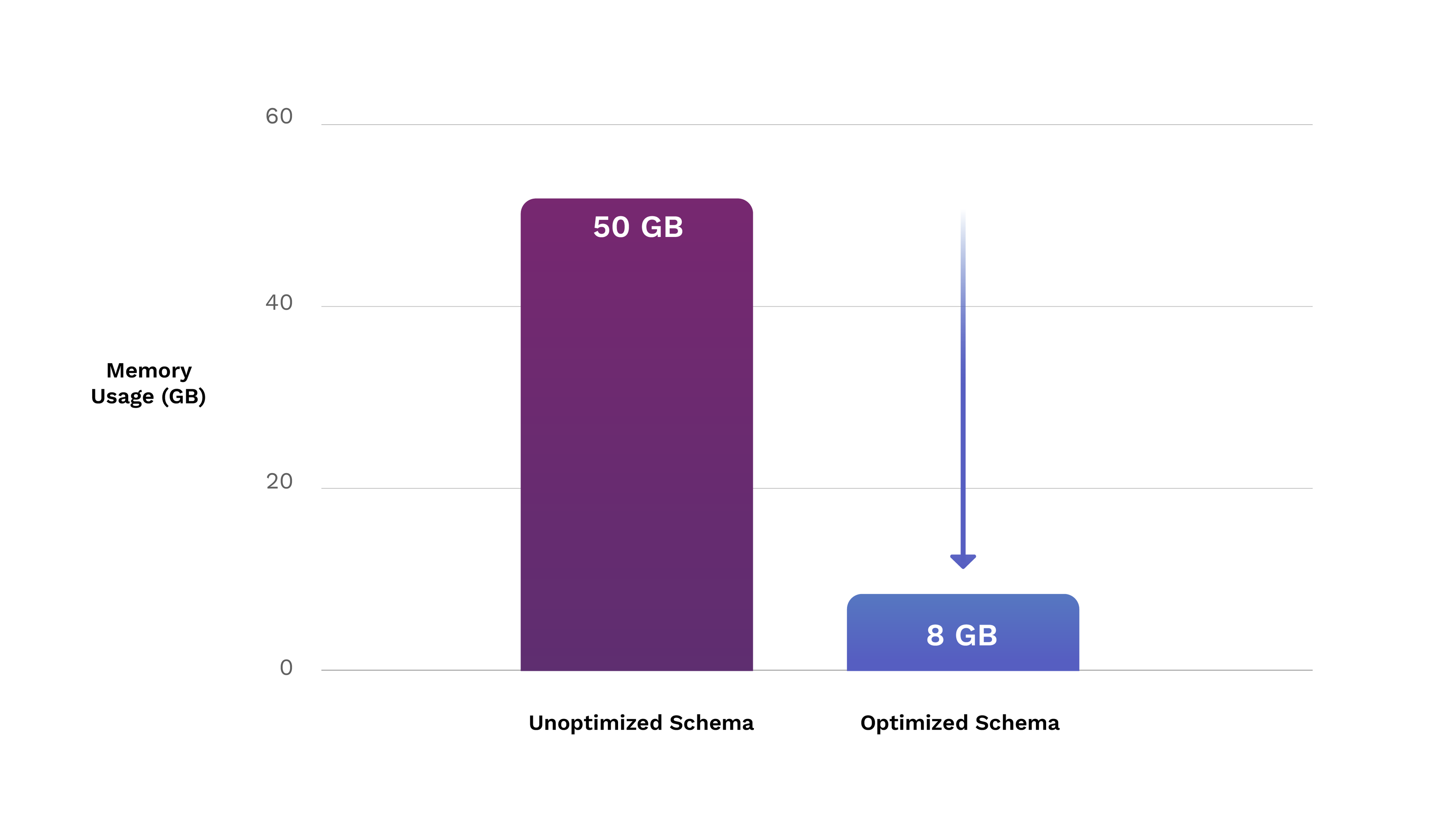
Out: 6347066223That translates to 6.3 GB — and including household data only adds 2 more GB. 6x better than without the tight schema! 8 GB isn’t only smaller, it can also fit into even personal machines much more neatly than 50 GB.
Can We Do Even Better?
It’s certainly worth noting that having expert knowledge could save time — for example, if there’s a general understanding of dataset requirements, our type inference can be used safely on only a subset of the data. This is out of the scope for this article (after all, we wanted to find a one-time cost that could yield improvement without care for time + resources).
Closing Out
A lot of data in this world is stored in sets of CSVs, with various methods of defining their schema — some easy to use, some less so. CSVs can be compressed, but, ultimately, are hardly made for this, and reading them can be tedious — in some systems, you have to define the type for each column to avoid safe-but-costly type inference! Parquet offers compression and schema definition alike. Here, we see what this can offer: both improved disk usage and the capability to bring some datasets in striking distance of smaller computers with minimal input from the user. That last part is what makes this process especially sweet: once it’s done, all the work is encoded in the Parquet file.
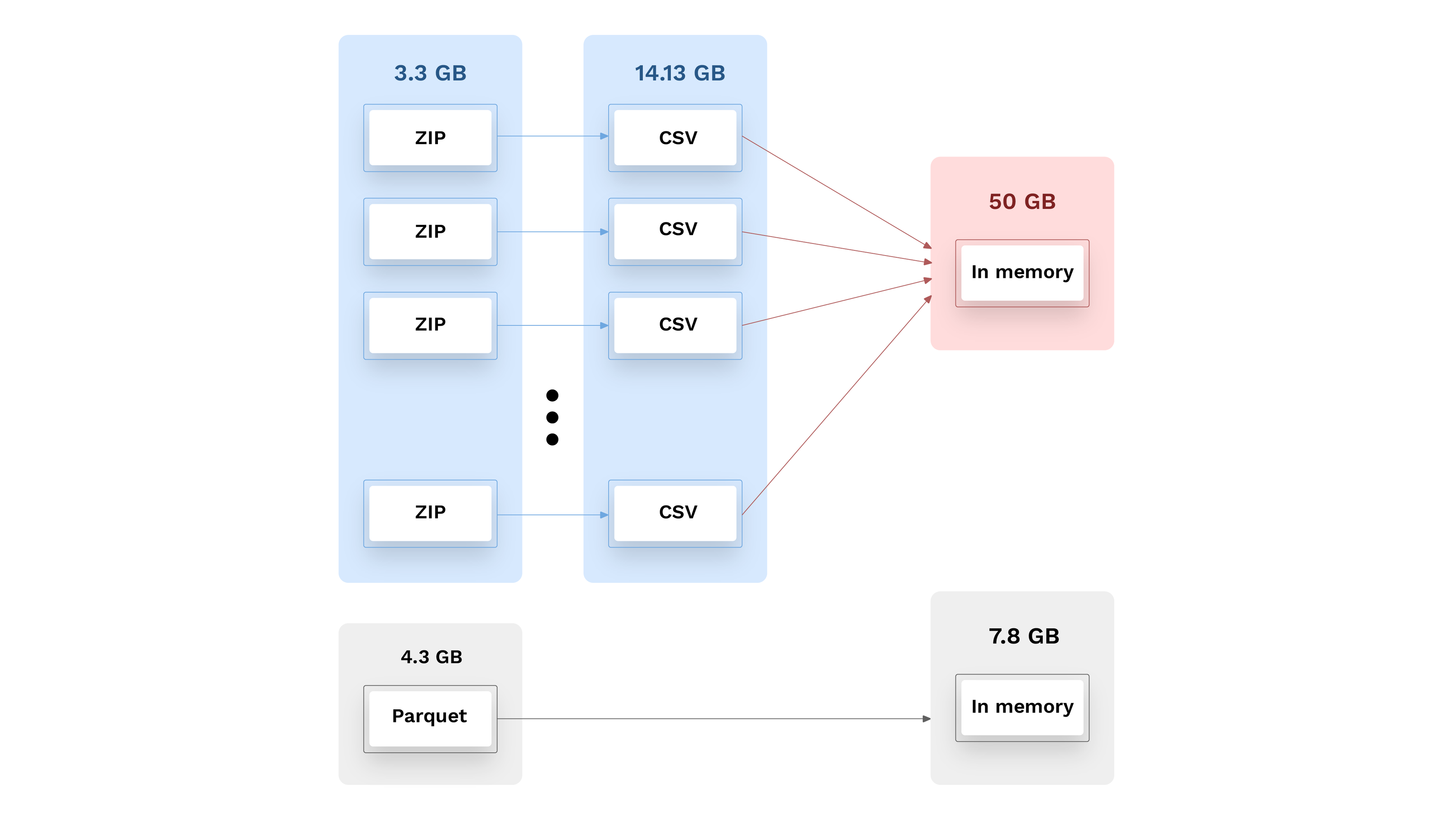
At this point, this file can be served and used as 4 GB on disk. While the Parquet zip is 1 GB larger than the original zip files, it is readable in this compressed form and comes as individual files. Even better are the in-memory benefits that require no effort from future users, offering 8GB in-memory instead of 50 GB.
This method, too, is portable in concept — all that’s necessary is tuning names and performing some explicit definition if errors arise. Of course, this ease is best attributed to Ibis — such easy access to analytics over all this data was made possible with it.
Then, tying this all together with its powerful utilities for memory and I/O, is PyArrow, which made it easy to take multiple CSVs and amass them into one dataset, never overwhelming system memory, and directing all back to that Parquet file.
Now, it may still be unclear why one would use Parquet instead of CSV here — maybe you simply have enough resources to work with all this data at once, or you’d load a subset to alleviate this without a complete migration. Each is as portable as the other, and you could read them anywhere — but that misses the nuance. Anywhere you read CSV, effort is needed to stage the data properly, and you must always pay the cost of parsing, setting the schema, and so on. Having Parquet immediately, without further work, enables use cases such as an app that uses it for backend data, or the ability to use Arrow FlightSQL, or passing to a tool that uses Parquet, such as XGBoost, or even SageMaker. This up-front investment to use Parquet provides the opportunity to spend future work on using your data, rather than reading it.
It’s a powerful data analytics and management stack, and scales well beyond a laptop. Tomorrow, we’re publishing another blog that showcases powerful tools - Apache Arrow, Ibis, and CuDF - to analyze this PUMS dataset. Stay tuned!
Photo by: Verne Ho
Previous Article
Keep up with us



